16| Phylogenetics in Ecology
Phylogenetic Systematics
The goal of phylogenetic systematics or "cladistics" is to construct classifications that accurately reflect phylogeny. To accomplish this end, states of numerous characters are determined and carefully analyzed for all species that are members of a given monophyletic group. Ancestral states do not provide any phylogenetic signal, but shared derived, or synapomorphic, character states do contain information that allows phylogenetic inference. Character states that arise independently (convergences) are misleading because they suggest relatedness where there is none. The challenge for phylogenetic systematists is to identify shared derived character states (synapomorphies) and to exclude convergent ones. Comparison with appropriate related "outgroups" allows systematists to identify ancestral traits and to "polarize" character state changes of probable shared derived characteristics. Parameter space is then searched for the most probable phylogenetic trees using various criteria, including distance, parsimony, and maximum liklihood. A subsampling method (bootstrapping) allows estimation of confidence in various branching patterns. With an adequate data set of character states, a phylogeny can be resolved and a most robust tree identified for use in further analyses. Using a closely related "outgroup," such a tree can be "rooted" to produce a "cladogram" that shows probable genealogical relationships among members of the group concerned. Past history can be recovered, at least to some extent, from current character states. Exploitation of these techniques to their fullest potential obviously requires examination of as many related taxa as possible -- extinctions of existing species truly become "lost pages" in the unread book of life.
Modern molecular techniques, particularly DNA amplification and sequencing, now allow systematists to reconstruct probable phylogenies of diverse monophyletic groups of organisms (Hillis et al. 1996). Any phylogeny is no more than a hypothesis, but the advent of molecular approaches combined with increasingly rigorous statistical procedures has greatly increased our confidence in particular "resolved" phylogenetic trees.
Armed with such a phylogeny showing the probable relationships among members of a monophyletic lineage, one can overlay various character states such as behavioral and ecological traits to examine the probable course of evolution. Powerful inferences can then be made about the sequence of changes that occurred during the evolution and adaptive radiation of that group. Modern comparative methods have also been developed that allow scientists to reconstruct probable ancestral states from those of surviving descendants. Obviously, we have made substantial progress in beginning to be able to decipher the vanishing book of life!
Vicariance Biogeography
As a consequence of the emerging awareness of continental drift, a vigorous new branch of biogeography has arisen. Prior attempts at explaining the current geographic distributions of organisms relied heavily on improbable movements and episodes of dispersal. But the recognition that landmasses themselves actually break apart and move (aquatic systems behave analogously) has enabled distributions to be understood in terms of such geographic or "vicariant" events as well as in terms of dispersal.
Concordances among cladograms for different plant and animal groups in the same general region suggest that common vicariant events underlie their phylogenies. Ultimately, such patterns can be related to features of the geographic history of the area concerned in an "area cladogram" (Figure 16.1). Indeed, vicariance biogeographers have actually suggested that certain physical geographic events must have occurred (for many years prior to the accumulation of extensive geological evidence, biogeographic distributions were the strongest evidence for continental drift!).
-
Figure 16.1. Continental movements, including the breakup of Pangaea and Gondwanaland
plus the collision of the Indian plate with southern Asia, are depicted in an "area cladogram."
Phylogeny and the Modern Comparative Method
Evolution of phenotypes (or species) in a range of different environments is analogous to the response of individuals in a nested sequence of experimental treatments (Harvey and Pagel 1991). The analogue to an experiment begins by subjecting a population of individuals to the same treatment at the base of a monophyletic tree (Figure 16.2). Subpopulations are divided and redivided sequentially after varying intervals of time. Between divisions, each subpopulation is subjected to a particular environmental "treatment" that differs from that experienced by other subpopulations.
Under this analogy, however, no record is kept of the "treatments" administered to various subpopulations. An estimate of the probable historical record of "treatments" and "responses" can be reconstructed, however, from appropriate information on extant lineages, using parsimony, as follows. We assume that past environments for a given lineage tend to be similar to present environments ("treatments"). Organisms tend to occupy habitats similar to those occupied by their ancestors.

-
Figure 16.2. A phylogenetic tree can be viewed as an analogue to a nested experiment. Lineages from nodes 2 and 3 terminating in crosses represent extinct fossils. At the basal node 1, a population is subdivided into two subpopulations, which are subjected to different environmental "treatments." At each subsequent node, subpopulations are redivided and subjected to new varying "treatments." Probable states at ancestral nodes can be estimated from states of extant species at the tips of the phylogenetic branches. [Adapted from Harvey and Pagel (1991).]
Extant populations contain information about their evolutionary history because the structure of the phylogenetic tree is known (Figure 16.2). For example, parsimony (minimal evolution) suggests that the ancestral subpopulation at node 4 is likely to have had condition 2 at both states because a majority of descendant subpopulations exhibit that condition. However, node 5 would be more likely to have exhibited condition 1 as judged from the prevalent condition among its descendants. The ancestral condition at the base of the phylogeny, node 1, could have been either condition 1 or condition 2 at both states. Note also that the states of extant subpopulations tend to covary across environments ("treatments"), making the condition of either state a good predictor of the condition of the other state. Phenotypes a, c, and d have probably undergone convergent evolution, and phenotype b has probably converged on phenotypes e, f, g, all of which appear to have retained the probable ancestral state 2, 2.
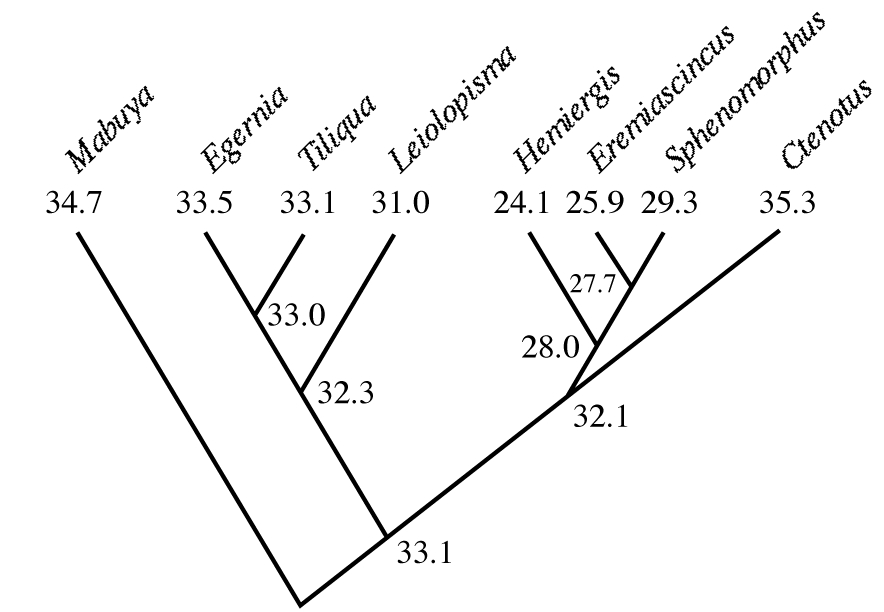
Probable ancestral states can be inferred from existing states of descendants using simple averages or in more complex ways involving maximum likelihood that minimize squared changes along all links of a phylogenetic tree. If branch lengths are known, such averages can be inversely weighted by the estimated distance between taxa. Although this approach is somewhat circular, it allows one to trace probable past evolutionary scenarios. An example for evolution of body temperatures among Australian skincid lizards is shown in Figure 16.3; this analysis suggests that the genus Ctenotus has evolved a higher active body temperature and that its relatives have evolved lower body temperatures.
-
Figure 16.3. Active body temperatures among seven genera of Australian skinks, with inferred ancestral body temperatures at various nodes. Note the high body temperature of Ctenotus and the lower body temperatures of its relatives Hemiergis, Eremiascincus, and Sphenomorphus. [Adapted from Huey and Bennett (1987).]
Phylogenetically Independent Contrasts
Until recently, ecologists have seldom considered how phylogenetic relatedness might affect observed patterns of covariation among phenotypic traits. Plotting various traits against one another, species-by-species is flawed because data points are not independent, strictly speaking. For example, if species exhibiting high values for both traits are closely related and those with lower values represent a distantly related clade of species that are themselves closely related, then an observed correlation is based on a sample size closer to two, rather than the total number of species. Hence the actual degrees of freedom are substantially less than would be the case if species were truly independent. A method of analysis that corrects for phylogenetic relatedness, known as independent contrasts, was suggested by Felsenstein (1985). This method compares the differences between related taxa, which are independent. Ancestral states can be estimated and used in independent contrasts as well. If a correlation between traits remains significant after applying these techniques to remove the phylogenetic effects of relatedness, then we have considerably greater confidence that the traits concerned covary as suspected.
Evolutionary Ecomorphology
Population biologists have long appreciated that an animal's morphology reflects its ecology. Consider desert lizards. Species that spend a lot of time in the open away from cover tend to have longer hind legs, relative to their snout-vent length, than those that stay closer to safe retreats. Longer legs increase running speed, hence facilitating use of open spaces. However, long-legged species move very clumsily through dense vegetation, demonstrating that there is actually a premium on shorter legs for species that exploit such closed-in microhabitats. Terrestrial species tend to have longer hind legs than arboreal species. Among sympatric lizards, limb proportions are correlated with their adaptations for climbing, burrowing, and other movements. Digging species typically possess larger front feet and more powerful forelegs than species that do not do much digging with their front limbs. Species that live entirely underground have reduced appendages or lack them altogether. Numerous other morphological correlates of the use of space exist. Arboreal lizards are typically long-tailed and slender, with claws or toe lamellae well suited for climbing. Indeed, number of toe lamellae, as well as their surface area, are intimately related to climbing ability. Among nocturnal geckos, terrestrial species tend to have larger eyes than arboreal species. Head proportions and jaw length are often reasonably accurate indicators of the size of a lizard's prey. Dentition sometimes reflects still more subtle aspects of diet, such as the agility and hardness of prey.
Such ecomorphological correlates may ultimately enable ecological predictions based on anatomical data. Morphological measures can often be estimated more easily and more objectively than ecological parameters. Morphometrics may represent average long-term responses to selection and hence reflect environmental conditions better than more direct measurements of the immediate ecological milieu.
Because anatomical parameters are usually much easier to estimate objectively than ecological ones, a variety of recent studies attempt to exploit such morphological correlates of ecology to make anatomical maps of ecological space and, in turn, to use these to analyze various aspects of community structure; these efforts have dealt with vertebrate taxa as divergent as bats, birds, fish, and lizards. In this approach, each species is represented as a point in an n-dimensional hypervolume whose coordinates are the morphological variates. These may be standardized as desired or log transformed. Euclidean distances between species are calculated as measures of dissimilarity. Distances from the centroid of the hypervolume can be exploited to judge the overall size of morphological space. If desired, dimensionality can be reduced and orthogonality achieved by a multivariate procedure such as principal components analysis (for an example, see Figure 16.4). (Euclidean distances between species remain unchanged when axes are rotated.) Spacing patterns between species, such as nearest neighbor distances, and other aspects of their position in morphological space can then be examined. The assumption is usually made that the arrangement of species in morphological space accurately reflects their ecological relationships, although this assumption is not easily verifiable and has seldom been directly tested.
-
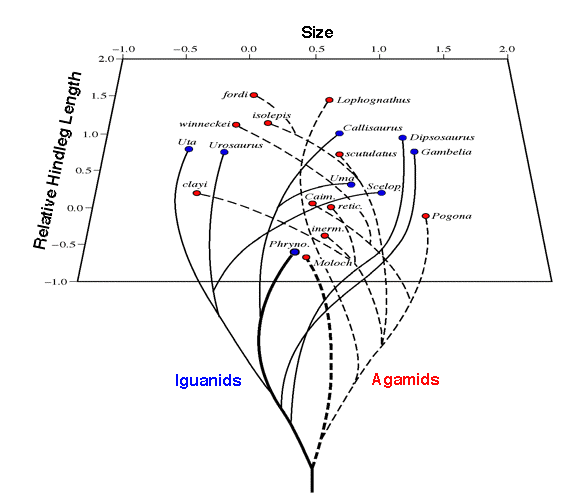
Figure 16.4. Positions of various North American and Australian lizard
species on the first two principal components of a 10-dimensional morphospace corresponding roughly
to body size and relative leg length, along with a hypothesis for phylogenetic relationships among
species. Note that the Australian thorny devil Moloch horridus and its American
"ecological equivalent," the desert horned lizard Phrynosoma platyrhinos,
are actually closer to one another than either is to another member of its own saurofauna.
[Key: Uta = Uta stansburiana, Uro = Urosaurus graciosus, Calli = Callisaurus draconoides,
Dipso = Dipsosaurus dorsalis, Crot = Crotaphytus wislizeni, Uma = Uma scoparia, Scelop =
Sceloporus magister, Phryno = Phrynosoma platyrhinos, Moloch = Moloch horridus,
inerm = Ctenophorus inermis, retic = Ctenophorus reticulatus, Caim = Caimanops
amphiboluroides, clayi = Ctenophorus clayi, winn = Diporiphora winneckei, isolepis = Ctenophorus
isolepis, fordi = Ctenophorus fordi, Loph = Lophognathus longirostris, scut = Ctenophorus
scutulatus, Pogona = Pogona minor.]
In one such study (Pianka 1986), ten morphometric variables were measured for each individual lizard collected: snout-vent and tail length, the length, width, and depth of the head, and the lengths of the jaw, forefoot, forearm, hindfoot, and hindleg. Even though sexual dimorphism occurs in some species, sexes were lumped for simplicity and ease of analysis. These measurements were strongly positively correlated with one another over all 92 species (mean correlation coefficient = 0.75). Average morphometric measures were used to represent each species as a point in a 10-dimensional morphospace. Each measurement was given equal weight by standardizing by subtraction of the mean value for all 92 species and division by the standard deviation across all species: this z transformation standardization procedure results in a mean score of 0.0 and a standard deviation of 1.0 for each variate. Distances from this standardized hypervolume's centroid (representing the overall "average" lizard species) were calculated for each species and averaged for each continental lizard fauna. Euclidean distances between all pairs of species were computed, and nearest neighbor distances identified for each. Various morphometric statistics were computed for each continent separately as well as for all intracontinental plus intercontinental pairs of species.
Anatomically, Kalahari lizard species are appreciably more similar than are North American or Australian lizard species. Both the overall average and nearest neighbor Euclidean distances are smaller and less variable in Kalahari lizards than on the other two continental-desert lizard systems. Euclidean distances between species are most variable in Australia, probably partially due to the larger number of species there. The overall volume of morphospace occupied by Kalahari lizards, as judged by distances from the centroid, is more compact than that occupied by lizards in North America
and Australia. To reduce dimensionality, a multivariate technique called principal components analysis was performed on these data. This method changes the coordinate system by identifying new orthogonal axes (principal components) that reduce residual variance. When all 92 species are considered together, the first three principal components reduce overall variance by 77.7 percent, 11.6 percent, and 6.7 percent (total 96 percent), respectively. Centroids for each of the continental faunas deviate from the overall centroid of all 92 species. Interestingly enough, even on these very crude morphological dimensions, the Australian thorny devil Moloch horridus and its American "ecological equivalent," the desert horned lizard Phrynosoma platyrhinos, are actually closer to one another than either is to another member of its own continent's lizard fauna. Both thorny devils and horned lizards are ant specialists.
Selected References
Phylogenetic Systematics
Eldredge and Cracraft (1980); Garland (1992); Garland et al. (1990, 1991, 1992); Harvey and Pagel (1991); Hennig (1966); Hillis et al. (1996); Ridley (1983); Rosen (1978, 1979); Swofford et al. (1996); Wiley (1981).
Vicariance Biogeography
Bermingham and Avise (1986); Nelson and Platnick (1981); Platnick and Nelson (1978).
Phylogeny and the Modern Comparative Method
Brooks and McLennan (1991); Felsenstein (1985, 1988); Garland (1991); Garland and Adolf (1994); Garland et al. (1991, 1992, 1999); Hansen and Martin (1996); Harvey and Pagel (1991); Harvey and Purvis (1991); Huey and Bennett (1987); Mayden (1992); Martins and Garland (1991); Martins and Hansen (1997); Pagel (1998); Ridley (1983); Schluter et al. (1997).
Phylogenetically Independent Contrasts
Felsenstein (1985, 1988); Harvey and Pagel (1991).
Evolutionary Ecomorphology
Ananjeva (1977); Bonine and Garland (1999); Cadle and Greene (1993); Collette (1961); Findley (1973, 1976); Gatz (1979a, 1979b); Grinnell (1924); Hecht (1952); Hespenheide (1973); Hotton (1955); Karr and James (1975); Lundelius (1957); Pianka (1986); Ricklefs and Travis (1980); Ricklefs et al. (1981); Werner (1968); Wainwright and Reilly (1994); Winemiller (1991).
|